40,000 women die from breast cancer annually in the US, and breast cancer is still the most common invasive cancer in women. 70-80% of these patients have estrogen receptor positive (ER+) disease which is currently treated with hormonal therapy, especially aromatase inhibitors (AIs) as first line therapy. Resistance to AIs is a major reason for disease recurrence and metastatic disease. AIs were designed specifically to inhibit aromatase (encoded by CYP19A1) activity, the enzyme that catalyzes the rate limiting step that converts androgens to estrogens. Estrogens are the major driving force for ER+ breast cancer. Among third generation AIs, letrozole and anastrozole are non-steroidal, while exemestane is steroidal. Prospective clinical trials have showed no differences in efficacy among these three AIs, perhaps due to the fact that results were always compared at the population level. However, our preliminary data indicate that knowledge of the host germline genetic background could help us to select AIs to treat specific individuals and to predict response to AI therapy. The purpose of this study is to understand how and why germline genetic variants might be associated with individual variation in in AI response. Understanding mechanisms for variation in AI response would help us design individualized therapies to treat ER+ breast cancer, especially for patients who are resistant to AI treatment
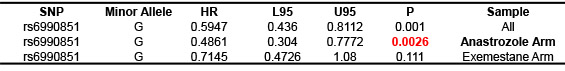
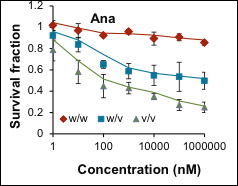
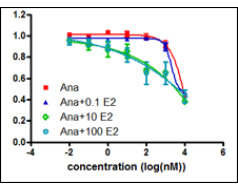
Since AIs block aromatase activity, suppression of estrogen formation is the pharmacodynamic effect of AIs. We set out to test whether genetic variation might contribute to AI efficacy through suppression of estrogen synthesis. To test that hypothesis, we took advantage of a series of large clinical trials for which we have extensive genomics and clinical response information. The first trial was conducted at Mayo Clinic, M.D. Anderson and MSK (Memorial Sloan Kettering) and the trial recruited over 800 post-menopausal women with primary early stage breast cancer treated with anastrozole. We have genome-wide SNP and hormone levels measured before and after patients had been treated with anastrozole for six months. We then performed a genome-wide association study (GWAS) for 624 of these patients for whom we had complete data to identify SNPs associated with estrogen level changes in women with ER+ breast cancer who were treated with anastrozole. Replication was then performed with disease recurrence as the phenotype for a second GWAS using samples from the MA.27 clinical trial with over 7500 patients randomized to anastrozole and exemestane treatment of post-menopausal women with ER+ breast cancer. That association study showed that a SNP in CSMD1 was associated with differential time to disease recurrence between the two AIs. That is, the variant genotype had a protective effect in the anastrozole arm only (Table 1). Functionally, the SNP influenced CSMD1 and, in turn, CYP19 gene expression, which contributed to SNP dependent response to anastrozole (Figure 1). Our data also suggested, surprisingly, that anastrozole might be an ERα ligand and function as ER degradator, especially in the presence of low dose E2. These findings have significant clinical implication: 1. The SNP can be used as a biomarker for selection of patients for anastrozole treatment. 2. The idea of combination of anastrozole and low dose estrogen to degrade ERα might be an alternative strategy to sensitize patients to anastrozole, especially in anastrozole resistant patients (Figure 2). 3. Our long term goal is to develop better therapeutic agents that can take advantage of our new findings.